Manufacturing human tissue from a patient’s own cells has long been the foundational promise of regenerative medicine. Two bottlenecks have kept that promise out of reach: the inability to manufacture induced pluripotent stem cells (iPSCs) with industrial consistency across diverse donors, and the inability to engineer those cells into functional tissue regardless of the patient’s genetic background. Here, we report the first end-to-end demonstration that both bottlenecks can be solved simultaneously and in combination. Using Cellino’s automated, AI-guided iPSC manufacturing platform and Polyphron’s autonomous tissue foundry, we manufactured four clonal iPSC lines from demographically diverse donors and engineered them into structurally defined cortical tissue constructs with equivalent alignment across all genetic backgrounds, without having to re-optimize the platform from scratch for each new donor.
Introduction
The advent of reprogramming technologies enabled the generation of iPSCs from various somatic cell types, providing the first pathway to manufacturing personalized human biology at scale. These cells hold immense promise across two distinct paradigms. First, as the foundation for advanced regenerative medicine, they enable tissue replacement therapies capable of completely reversing disease pathology rather than merely managing symptoms. Second, they unlock the creation of population-level models of human tissue that capture patient-specific biology, providing a high-fidelity validation substrate that can close the loop for emerging applications of artificial intelligence in biology, such as in silico drug design, toxicology, screening, and even simulation of early stage clinical trials at a throughput and genetic diversity that no animal model or immortalized cell line can match.
Early optimism suggested that iPSCs would serve as a standardized commodity, with uniform differentiation capacities similar to those of human embryonic stem cells. In practice, the field has encountered fundamental biological limits. Multiple studies have revealed profound heterogeneity in the differentiation potential of iPSC lines, a crucial variability that severely undermines the efficiency and scalability of existing approaches. This variability arises from multiple sources, including the patient’s genetic background, sex, age, the epigenetic memory of the tissue of origin, and the mechanics of the reprogramming methods.
To unlock curative therapies and scale the manufacturing of predictive tissues, two problems must be solved simultaneously. First, patient-derived iPSCs must be manufactured with industrial consistency across a diverse population. Second, those cells must be engineered into functional tissues whose architecture meets defined tolerances, regardless of the patient’s genetic background. Solving either problem in isolation is insufficient; a perfectly manufactured iPSC line that cannot be reliably differentiated into functional tissue has no clinical or commercial utility, and a powerful tissue engineering platform that fails across genetically diverse lines cannot target nor model a real patient population.
This white paper reports results from a collaboration between Cellino and Polyphron addressing both bottlenecks end-to-end. Cellino manufactured four clonal iPSC lines from demographically diverse donors using its automated optical bioprocess. Polyphron evaluated these lines through its tissue foundry, demonstrating reproducible differentiation and structure across all donor backgrounds via a cross-donor optimization framework that identified line-specific perturbations without re-optimizing from scratch for each new genotype, demonstrating a path toward scalable, genotype-matched tissue manufacturing.
Automated iPSC Manufacturing
Today, iPSC generation relies on manual processes, highly skilled operators for in-process decision-making, and dedicated cleanrooms for each patient, a model that does not scale to serve a diverse and growing patient population. Cellino’s automated closed-compatible biomanufacturing technology, based on a proprietary optical bioprocess, overcomes these constraints. The technology employs automated imaging at frequent intervals, artificial intelligence (AI)-based image analysis for in-process assessment, and laser-based in situ removal of cells with single-cell precision, enabling passage-free culture and continuous process control. AI-driven image analysis replaces subjective operator assessments with consistent phenotypic characterization throughout every stage of iPSC generation. The optical bioprocess is optimal for closed manufacturing and supports small-footprint, parallelized processing that reduces contamination risk, minimizes operator-to-operator variability, and enables scalable production of iPSCs across a broad and demographically diverse population. Cellino’s iPSC manufacturing technology received the FDA’s Advanced Manufacturing Technology (AMT) designation.
For this collaboration, Cellino manufactured four clonal iPSC lines from demographically diverse donors using its proprietary optical bioprocess (Table 1). CD34+ cells isolated from non-mobilized peripheral blood were reprogrammed using Sendai virus and cultured on Cellino’s platform, which provides automated incubation, imaging, media exchange, and laser-based cell removal. Following reprogramming, colonies were automatically identified and tracked over time, using Cellino’s AI-based image analysis algorithms to extract morphological features from label-free brightfield images. Cellino’s AI models are trained using supervised deep learning on labeled datasets, pairing label-free brightfield image stacks with fluorescence-derived ground truth maps. The ground truth is generated from a combination of fluorescent cell reporter lines, live-cell staining, and expert human annotation of brightfield images from datasets that span multiple iPSC clones and culture conditions representative of the optical bioprocess. Because colonies are continuously observed throughout the manufacturing lifecycle, these image-derived measurements form time-resolved phenotypic profiles that capture dynamic properties, such as growth rate, and offer a rich feature space enabling a more informed clone selection process. Using this data, final clones were selected for expansion and banking, employing laser-based cell removal to eliminate unwanted cells and colonies from the growth surface. All four lines were characterized to ensure they met the acceptance criteria for a range of critical quality attributes, including sterility, residual Sendai vector clearance, a normal G-band karyotype, expression of intracellular and extracellular pluripotency-associated markers, and the ability to differentiate into all three germ layers. This end-to-end process, from reprogramming through AI-guided clone selection to fully characterized, cryopreserved banks, demonstrates a scalable, automated manufacturing workflow in which AI-driven decision-making assists operators at critical steps, laying the groundwork for distributed, autonomous production of patient-specific iPSCs.

Table 1. iPSC lines manufactured by Cellino. Four clonal lines from demographically diverse donors, spanning both sexes, ages 24–54, and three ethnic backgrounds, all reprogrammed from peripheral blood using the CytoTune Sendai kit.
The availability of four well-characterized, quality-controlled iPSC lines from demographically diverse donors provided the ideal substrate to stress-test the second bottleneck: whether a tissue engineering platform can produce structurally equivalent outcomes independent of donor genetic background.
Evaluating Polyphron’s Platform Across Donor Lines
Polyphron has previously demonstrated computational control of a key architectural feature of human cortical tissue, axonal-dendritic alignment of excitatory neurons, achieving near-native directional organization (i.e., 90°) in 3D neuronal cultures using its tissue foundry (more here).
To prove the foundry’s capability to produce structurally similar tissue across different donors, differentiated admixtures of inhibitory and excitatory cortical neurons derived from Cellino’s four iPSC lines were cultured in 3D microfluidic devices and perturbed with a shared set of developmentally-inspired morphogenetic perturbations. Neurite alignment was monitored longitudinally and quantified using Polyphron’s non-destructive AI phenotyping approach, and the results were benchmarked against those obtained in the previous study. To stress-test the platform's ability to generalize from data-rich to data-sparse donors, mimicking the real-world scenario of onboarding a new patient line with limited experimental history, the initial experimental space was intentionally designed with asymmetric data coverage. Donors 3 and 4 were comprehensively mapped, with temporal trajectories measured across 36 conditions each. Donors 1 and 2 were perturbed with fewer conditions and monitored for a fraction of the time (Table 2).

Table 2. Experimental coverage per donor iPSC line at the time of prediction, prior to validation plate design. Asymmetric coverage was intentional, designed to stress-test the trajectory transfer framework's ability to generalize from data-rich to data-sparse donors.
The resulting phenotypes confirmed what has been previously observed in the field when applying the same protocol across different iPS lines. Inter-donor variability, here quantified as the divergence in alignment across different lines when perturbed with the same cocktail, was apparent, resulting in up to a 9-fold variance in alignment across shared perturbation conditions. This confirmed that a universal, static perturbation set is not sufficient for tissue manufacturing at scale (Figure 1).
Traditional approaches have historically tackled the problem by adapting experimental protocols through exhaustive, often manual trial-and-error screens for each new patient line, rendering the process intractable and uncommercially viable at scale. Conversely, Polyphron’s platform exploited the shared perturbation space and donor-specific trajectories to quantify inter-donor differences and transfer trajectories from data-rich lines (Donors 3 and 4) to the data-sparse lines (Donors 1 and 2), using the target line's initial alignment values as a localized anchor. Integrating information from perturbations and temporal dynamics generated a composite trajectory score, leading to the identification of 5 perturbations, new or existing, predicted to drive alignment for each specific genotype, without requiring that genotype to have been fully characterized in advance.
Upon prospective validation, predictions were confirmed with all four lines reaching alignment values close to 90°. The framework correctly identified top-performing perturbations for every donor and even identified morphogenetic conditions that the lines had never previously encountered. Across all four donors, 19 of 20 top-performing wells effectively tied the alignment values seen for the optimized reference line (Figure 1).
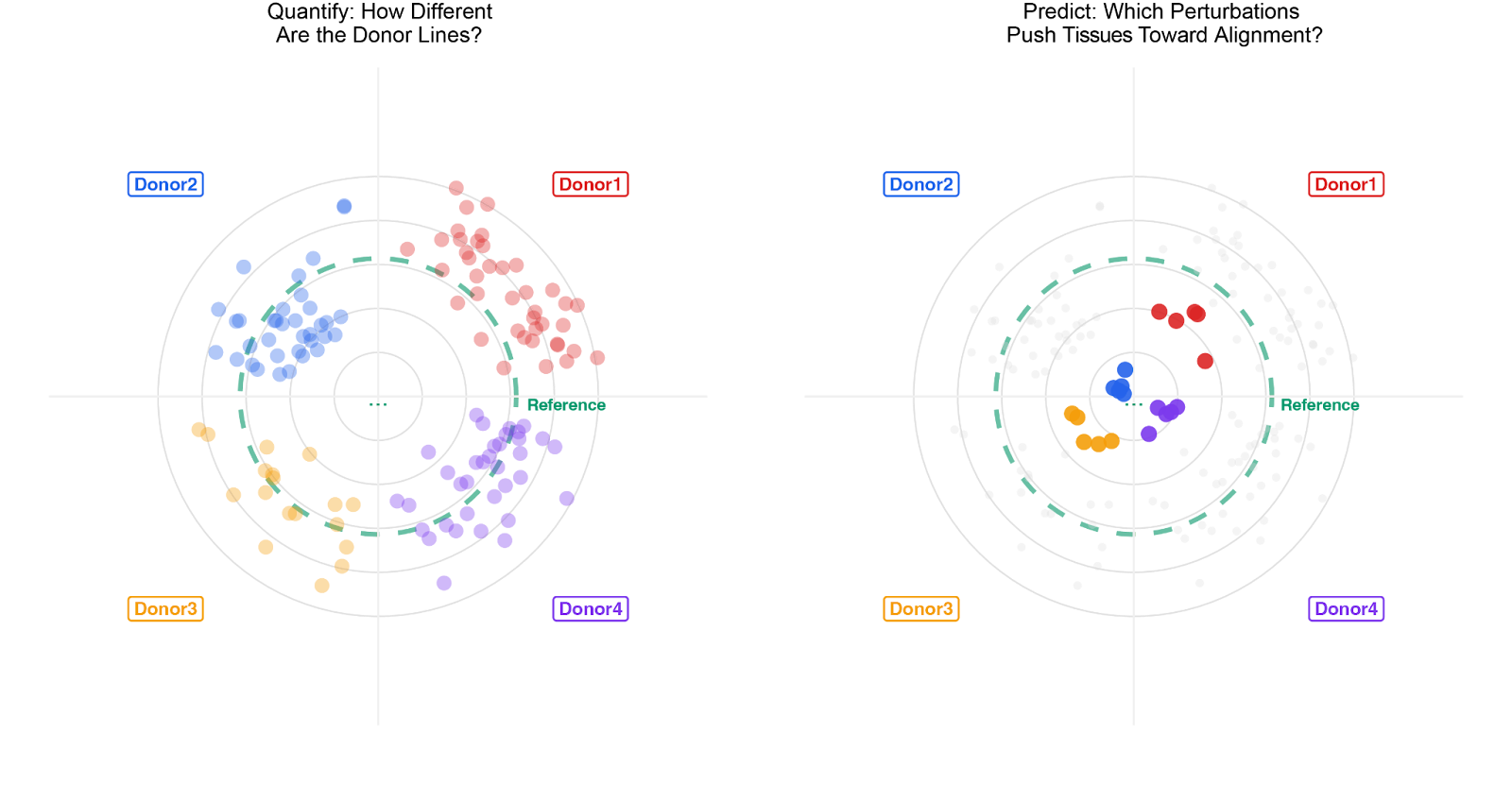
Figure 1: Cross-donor generalization of cortical alignment through AI-guided optimization. Left: bull's-eye plots quantifying inter-donor phenotypic variability across four iPSC lines (Donors 1–4) and a reference line. Each dot represents a single tissue unit; position toward the center indicates higher axonal-dendritic alignment. Initial perturbation conditions produce significantly different structural phenotypes across donors. Right: after quantification, the platform identifies perturbations that pushed all lines toward highly aligned tissue
By mapping donor-specific trajectories, the platform successfully identified perturbations that engineered stable, highly aligned tissue across every genetic background tested (Figure 2). The reference line required three active learning iterations and thousands of observations to converge on its optimal perturbation set. By learning from inter-donor differences and line-specific trajectories, the platform reduced the experimental footprint by 12-fold for newly onboarded donors.
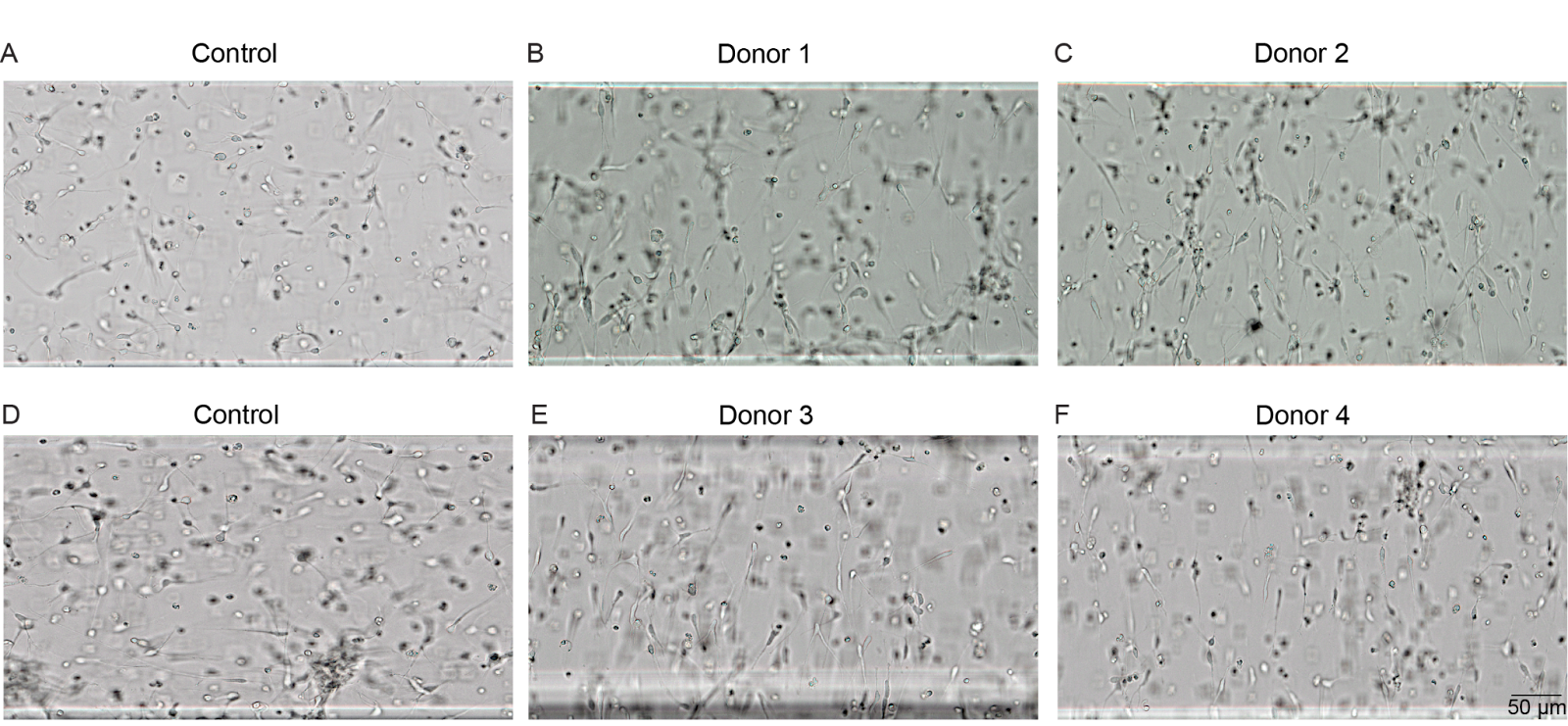

Figure 2: Reproducible neurite alignment across donor iPSC lines. Bright-field images of cortical neuron constructs in Polyphron’s platform. (A, D) Control conditions showing randomly oriented neurites. (B) Donor 1, (C) Donor 2, (E) Donor 3, and (F) Donor 4 under donor-specific optimized perturbation conditions, demonstrating consistent directional alignment across all four genetically distinct donor lines. Scale bar, 50 µm.
Molecular Characterization of the iPSC lines
To establish an independent molecular framework for interpreting cross-donor heterogeneity, bulk RNA sequencing was performed on all four iPSC lines at the pluripotent stage (4 biological replicates per line, 13,912 protein-coding genes after filtering), and single-cell RNA sequencing was performed on the differentiated cortical cultures (33,219 cells post-QC). These molecular profiles were generated in parallel to retrospectively validate the biological drivers of divergence, rather than serving as prior inputs for the foundry’s predictions.
Transcriptomic profiling of iPSCs revealed structured baseline divergence, consistent with the known transcriptomic heterogeneity among iPSC lines from different donors. Principal Component Analysis and differential expression analysis revealed that this heterogeneity distributes along two axes of biological variation, each with a distinct molecular identity and functional implication.
- Axis 1: Epigenetic repression (PC1, 35.7% variance). The dominant axis of transcriptional variation isolates Donor 2 from all other lines. Targeted analysis of known epigenetic regulators across multiple functional categories identified the molecular basis for this divergence. Donor 2 showed reduced expression of genes associated with genome activation and occupancy, suggesting a distinctly repressive pre-differentiation state and potentially reduced accessibility at neuronal-specific genomic regions.
- Axis 2: Progenitor-to-neuronal commitment (PC2, 17.2% variance). The second axis separated Donor 1 from Donor 3, with Donor 4 falling in the middle. Donor 1 expressed progenitor- and stem cell-like transcriptional programs. In contrast, Donor 3 expressed a neurogenically primed signature and neuron-specific epigenetic drivers.
Single-cell RNA sequencing on the differentiated cultures provided direct functional confirmation that these pre-differentiation axes have measurable consequences. After cell-type annotation using marker scoring and reference-based annotation, the four lines produced dramatically different neuronal populations. Donor 1, the most progenitor-like line by bulk transcriptomics, retained the highest progenitor fraction after differentiation (69.3% Progenitor/RG). Donor 2, the chromatin-repressed outlier, produced a unique inhibitory neuron bias (44.7% InN). Overall, these findings confirmed that inter-donor transcriptomic heterogeneity at the iPSC stage is a biologically meaningful signal with direct consequences for differentiation outcomes.
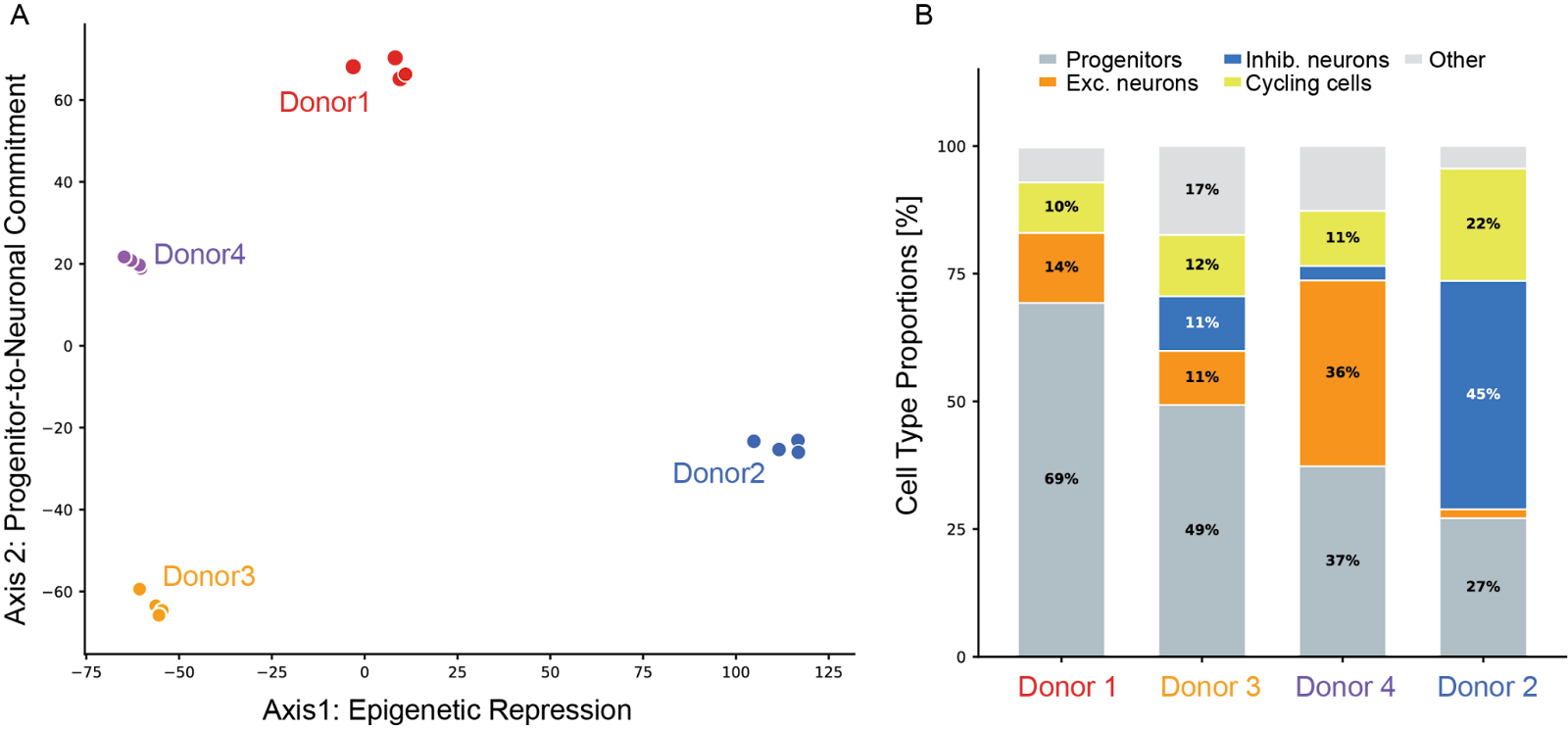
Figure 3: Molecular characterization of the iPSC lines. (A) Principal Component Analysis of bulk RNA-seq from four iPSC lines at the pluripotent stage (4 replicates per line, 13,912 protein-coding genes). Axis separates Donor 2 along an epigenetic repression axis; Axis 2 separates Donor 1 from Donor 3 along a progenitor-to-neuronal commitment axis. (B) Cell-type proportions from single-cell RNA-seq of differentiated cortical cultures.
Engineering Around Biological Baselines
The platform's task was to engineer equivalent structural outcomes from these highly divergent starting points. Pairwise transcriptomic distances at the iPSC stage were not statistically linked to pairwise alignment distances post-differentiation, demonstrating that Polyphron’s platform can accommodate molecularly divergent lines and lead them to structurally similar phenotypes under optimized conditions. Notably, both Donor 3, the most neurally committed line, and Donor 2, the most transcriptomically divergent line with the most repressive chromatin state, achieved the highest alignment, the two lines furthest from a common biological baseline converging on the same structural target.
The practical consequence is that onboarding a new donor line does not require prior knowledge of its chromatin state, its position on the commitment axis, or its expected cell-type proportions. By quantifying the starting heterogeneity, Polyphron’s foundry empirically identifies the precise morphogenetic perturbations required to override a donor's default trajectory and force convergence on the target tissue architecture. By doing so, it drastically reduces the number of experiments traditionally required to manufacture human tissues at scale.
Conclusion
These results signal a fundamental shift in the economics and scalability of iPSC-based tissue manufacturing. Platforms that rely on static protocols will inevitably fail across diverse patient populations. Automated, reproducible iPSC manufacturing, as demonstrated by Cellino's optical bioprocess, is a prerequisite for making this signal mappable. Without manufacturing consistency, biological variability and technical variability are indistinguishable.
We have previously demonstrated the capacity of this foundry to seamlessly cross tissue contexts (e.g., cortical to cardiac). We have now validated its ability to achieve multi-donor generalization within a single tissue context, requiring zero architectural changes to the underlying platform. As our foundry scales and generates tissues from diverse donors and backgrounds, it will improve its ability to leverage observed trajectories and generalize to new lines, reducing the experimental footprint and the optimization burden from hundreds of runs to tens.
Together, Cellino and Polyphron are building the closed-loop manufacturing infrastructure required to make functional human tissue fully specifiable, reproducible, and scalable at the population level, treating genetic diversity as a programmable variable rather than an absolute limit.
